libcbm run of tutorial 2 from CBM-CFS3¶
[1]:
import os, json
import pandas as pd
%matplotlib inline
Import the required packages from libcbm
- sit_cbm_factory: a module for initializing the CBM model from the CBM Standard import tool format
- cbm_simulator: simulates the sit dataset using the CBM model
- libcbm.resources: gets files for tutorial 2 that are bundled in libcbm
[2]:
from libcbm.input.sit import sit_cbm_factory
from libcbm.model.cbm import cbm_simulator
from libcbm.model.cbm.cbm_output import CBMOutput
from libcbm import resources
Setup¶
Load the standard import tool configuration at the specified path. This configuration encompasses the data source for the various sit inputs (sit_inventory, sit_classifiers etc.) and also the relationships of classifiers, and disturbance types to the default CBM parameters.
[3]:
config_path = os.path.join(
resources.get_test_resources_dir(), "cbm3_tutorial2", "sit_config.json"
)
sit = sit_cbm_factory.load_sit(config_path)
Initialize and validate the inventory in the sit dataset
[4]:
classifiers, inventory = sit_cbm_factory.initialize_inventory(sit)
Create storage for CBM simulation results. This is a built in method, and it is not mandatory since it is possible to interact with the simulation dataframes directly as well.
[5]:
cbm_output = CBMOutput(
classifier_map=sit.classifier_value_names,
disturbance_type_map=sit.disturbance_name_map,
)
Simulation¶
The following line of code spins up the CBM inventory and runs it through 100 timesteps.
[6]:
with sit_cbm_factory.initialize_cbm(sit) as cbm:
# Create a function to apply rule based disturbance events and transition rules based on the SIT input
rule_based_processor = sit_cbm_factory.create_sit_rule_based_processor(
sit, cbm
)
# The following line of code spins up the CBM inventory and runs it through 200 timesteps.
cbm_simulator.simulate(
cbm,
n_steps=200,
classifiers=classifiers,
inventory=inventory,
pre_dynamics_func=rule_based_processor.pre_dynamics_func,
reporting_func=cbm_output.append_simulation_result,
)
Pool Results¶
[7]:
pi = cbm_output.classifiers.to_pandas().merge(
cbm_output.pools.to_pandas(),
left_on=["identifier", "timestep"],
right_on=["identifier", "timestep"],
)
[8]:
pi.head()
[8]:
| identifier | timestep | Working_Species_Or_Leading_Species | Site_Quality | Density_Class | Working_Status | Input | SoftwoodMerch | SoftwoodFoliage | SoftwoodOther | ... | BelowGroundSlowSoil | SoftwoodStemSnag | SoftwoodBranchSnag | HardwoodStemSnag | HardwoodBranchSnag | CO2 | CH4 | CO | NO2 | Products | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 0 | BF | GOOD | D1 | W | 200.0 | 0.000000 | 0.000000 | 0.0 | ... | 10529.242115 | 5568.605479 | 1544.809522 | 0.0 | 0.0 | 690032.809697 | 631.259013 | 5681.261733 | 0.0 | 0.0 |
| 1 | 2 | 0 | BF | GOOD | D1 | W | 100.0 | 0.003124 | 0.376897 | 0.0 | ... | 5273.531854 | 2666.157167 | 666.417417 | 0.0 | 0.0 | 345255.294728 | 315.629506 | 2840.630866 | 0.0 | 0.0 |
| 2 | 3 | 0 | BF | GOOD | D1 | W | 100.0 | 0.030782 | 1.867345 | 0.0 | ... | 5278.708266 | 2553.024910 | 574.973378 | 0.0 | 0.0 | 345471.445778 | 315.629506 | 2840.630866 | 0.0 | 0.0 |
| 3 | 4 | 0 | BF | GOOD | D1 | W | 100.0 | 0.117354 | 4.744316 | 0.0 | ... | 5281.125622 | 2444.693447 | 496.077048 | 0.0 | 0.0 | 345669.654559 | 315.629506 | 2840.630866 | 0.0 | 0.0 |
| 4 | 5 | 0 | BF | GOOD | D1 | W | 100.0 | 0.303283 | 9.168135 | 0.0 | ... | 5281.488695 | 2340.959458 | 428.006664 | 0.0 | 0.0 | 345853.464520 | 315.629506 | 2840.630866 | 0.0 | 0.0 |
5 rows × 33 columns
[9]:
biomass_pools = [
"SoftwoodMerch",
"SoftwoodFoliage",
"SoftwoodOther",
"SoftwoodCoarseRoots",
"SoftwoodFineRoots",
"HardwoodMerch",
"HardwoodFoliage",
"HardwoodOther",
"HardwoodCoarseRoots",
"HardwoodFineRoots",
]
dom_pools = [
"AboveGroundVeryFastSoil",
"BelowGroundVeryFastSoil",
"AboveGroundFastSoil",
"BelowGroundFastSoil",
"MediumSoil",
"AboveGroundSlowSoil",
"BelowGroundSlowSoil",
"SoftwoodStemSnag",
"SoftwoodBranchSnag",
"HardwoodStemSnag",
"HardwoodBranchSnag",
]
biomass_result = pi[["timestep"] + biomass_pools]
dom_result = pi[["timestep"] + dom_pools]
total_eco_result = pi[["timestep"] + biomass_pools + dom_pools]
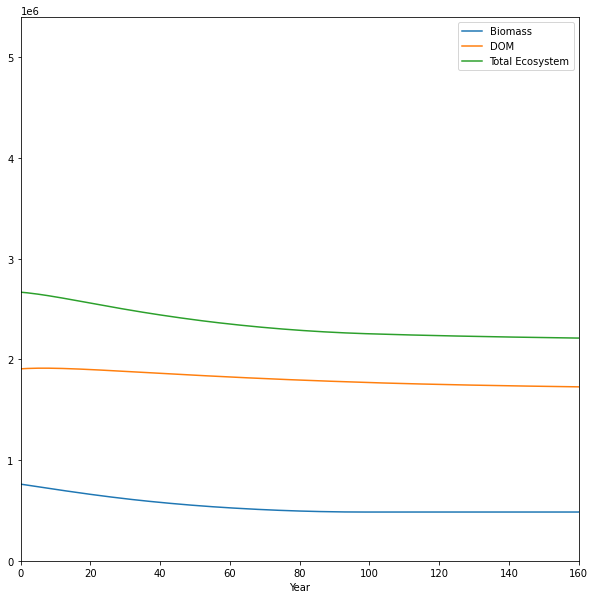
annual_carbon_stocks = pd.DataFrame(
{
"Year": pi["timestep"],
"Biomass": pi[biomass_pools].sum(axis=1),
"DOM": pi[dom_pools].sum(axis=1),
"Total Ecosystem": pi[biomass_pools + dom_pools].sum(axis=1),
}
)
annual_carbon_stocks.groupby("Year").sum().plot(
figsize=(10, 10), xlim=(0, 160), ylim=(0, 5.4e6)
)
[9]:
<Axes: xlabel='Year'>
State Variable Results¶
[10]:
si = cbm_output.state.to_pandas()
[11]:
si.head()
[11]:
| identifier | timestep | last_disturbance_type | last_disturbance_event | time_since_last_disturbance | time_since_land_class_change | growth_enabled | enabled | land_class | age | growth_multiplier | regeneration_delay | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 0 | Natural forest fire | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 1.0 | 0 |
| 1 | 2 | 0 | Natural forest fire | 0 | 1 | 1 | 1 | 1 | 0 | 1 | 1.0 | 0 |
| 2 | 3 | 0 | Natural forest fire | 0 | 2 | 2 | 1 | 1 | 0 | 2 | 1.0 | 0 |
| 3 | 4 | 0 | Natural forest fire | 0 | 3 | 3 | 1 | 1 | 0 | 3 | 1.0 | 0 |
| 4 | 5 | 0 | Natural forest fire | 0 | 4 | 4 | 1 | 1 | 0 | 4 | 1.0 | 0 |
[12]:
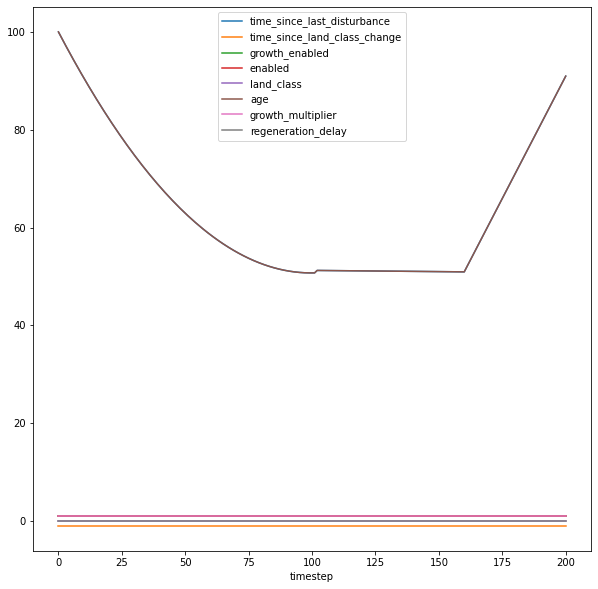
state_variables = [
"timestep",
"time_since_last_disturbance",
"time_since_land_class_change",
"growth_enabled",
"enabled",
"land_class",
"age",
"growth_multiplier",
"regeneration_delay",
]
si[state_variables].groupby("timestep").mean().plot(figsize=(10, 10))
[12]:
<Axes: xlabel='timestep'>
Flux Indicators¶
[13]:
fi = cbm_output.flux.to_pandas()
[14]:
fi.head()
[14]:
| identifier | timestep | DisturbanceCO2Production | DisturbanceCH4Production | DisturbanceCOProduction | DisturbanceBioCO2Emission | DisturbanceBioCH4Emission | DisturbanceBioCOEmission | DecayDOMCO2Emission | DisturbanceSoftProduction | ... | DisturbanceVFastBGToAir | DisturbanceFastAGToAir | DisturbanceFastBGToAir | DisturbanceMediumToAir | DisturbanceSlowAGToAir | DisturbanceSlowBGToAir | DisturbanceSWStemSnagToAir | DisturbanceSWBranchSnagToAir | DisturbanceHWStemSnagToAir | DisturbanceHWBranchSnagToAir | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 1 | 2 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 2 | 3 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 3 | 4 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 4 | 5 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
5 rows × 54 columns
[15]:
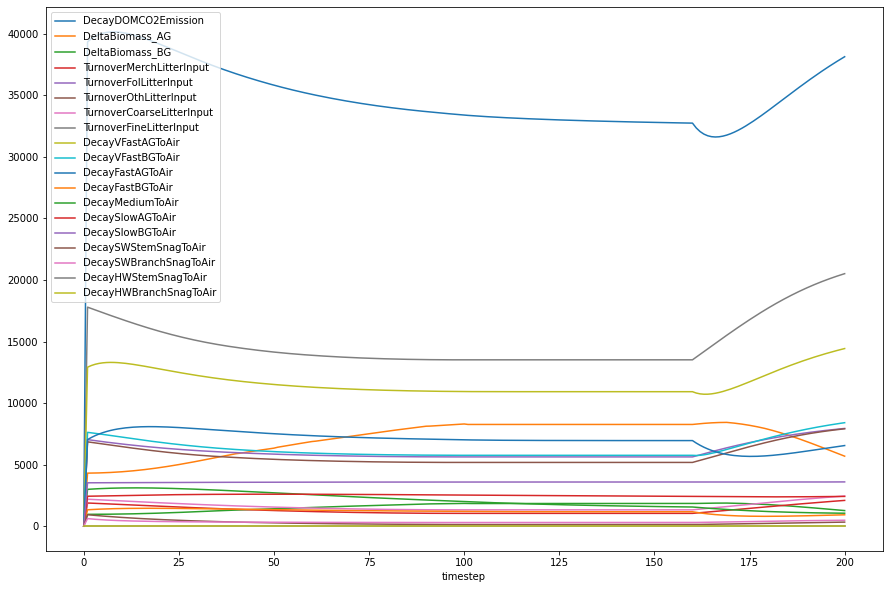
annual_process_fluxes = [
"DecayDOMCO2Emission",
"DeltaBiomass_AG",
"DeltaBiomass_BG",
"TurnoverMerchLitterInput",
"TurnoverFolLitterInput",
"TurnoverOthLitterInput",
"TurnoverCoarseLitterInput",
"TurnoverFineLitterInput",
"DecayVFastAGToAir",
"DecayVFastBGToAir",
"DecayFastAGToAir",
"DecayFastBGToAir",
"DecayMediumToAir",
"DecaySlowAGToAir",
"DecaySlowBGToAir",
"DecaySWStemSnagToAir",
"DecaySWBranchSnagToAir",
"DecayHWStemSnagToAir",
"DecayHWBranchSnagToAir",
]
[16]:
fi[["timestep"] + annual_process_fluxes].groupby("timestep").sum().plot(
figsize=(15, 10)
)
[16]:
<Axes: xlabel='timestep'>
Appendix¶
SIT source data¶
[17]:
sit.sit_data.age_classes
[17]:
| name | class_size | start_year | end_year | |
|---|---|---|---|---|
| 0 | AGEID0 | 0 | 0 | 0 |
| 1 | AGEID1 | 10 | 1 | 10 |
| 2 | AGEID2 | 10 | 11 | 20 |
| 3 | AGEID3 | 10 | 21 | 30 |
| 4 | AGEID4 | 10 | 31 | 40 |
| 5 | AGEID5 | 10 | 41 | 50 |
| 6 | AGEID6 | 10 | 51 | 60 |
| 7 | AGEID7 | 10 | 61 | 70 |
| 8 | AGEID8 | 10 | 71 | 80 |
| 9 | AGEID9 | 10 | 81 | 90 |
| 10 | AGEID10 | 10 | 91 | 100 |
| 11 | AGEID11 | 10 | 101 | 110 |
| 12 | AGEID12 | 10 | 111 | 120 |
| 13 | AGEID13 | 10 | 121 | 130 |
| 14 | AGEID14 | 10 | 131 | 140 |
| 15 | AGEID15 | 10 | 141 | 150 |
| 16 | AGEID16 | 10 | 151 | 160 |
| 17 | AGEID17 | 10 | 161 | 170 |
| 18 | AGEID18 | 10 | 171 | 180 |
| 19 | AGEID19 | 10 | 181 | 190 |
| 20 | AGEID20 | 10 | 191 | 200 |
[18]:
sit.sit_data.inventory
[18]:
| Working_Species_Or_Leading_Species | Site_Quality | Density_Class | Working_Status | age | area | delay | land_class | historical_disturbance_type | last_pass_disturbance_type | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | BF | GOOD | D1 | W | 0 | 200.0 | 0 | 0 | DISTID1 | DISTID1 |
| 1 | BF | GOOD | D1 | W | 1 | 100.0 | 0 | 0 | DISTID1 | DISTID1 |
| 2 | BF | GOOD | D1 | W | 2 | 100.0 | 0 | 0 | DISTID1 | DISTID1 |
| 3 | BF | GOOD | D1 | W | 3 | 100.0 | 0 | 0 | DISTID1 | DISTID1 |
| 4 | BF | GOOD | D1 | W | 4 | 100.0 | 0 | 0 | DISTID1 | DISTID1 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 196 | BF | GOOD | D1 | W | 196 | 100.0 | 0 | 0 | DISTID1 | DISTID1 |
| 197 | BF | GOOD | D1 | W | 197 | 100.0 | 0 | 0 | DISTID1 | DISTID1 |
| 198 | BF | GOOD | D1 | W | 198 | 100.0 | 0 | 0 | DISTID1 | DISTID1 |
| 199 | BF | GOOD | D1 | W | 199 | 100.0 | 0 | 0 | DISTID1 | DISTID1 |
| 200 | BF | GOOD | D1 | W | 200 | 100.0 | 0 | 0 | DISTID1 | DISTID1 |
201 rows × 10 columns
[19]:
sit.sit_data.classifiers
[19]:
| id | name | |
|---|---|---|
| 0 | 1 | Working_Species_Or_Leading_Species |
| 1 | 2 | Site_Quality |
| 2 | 3 | Density_Class |
| 3 | 4 | Working_Status |
[20]:
sit.sit_data.classifier_values
[20]:
| classifier_id | name | description | |
|---|---|---|---|
| 1 | 1 | BF | Balsam Fir |
| 2 | 1 | BS | Black Spruce |
| 3 | 1 | JL | Western Larch |
| 4 | 1 | JP | Jack Pine |
| 5 | 1 | OS | Other Spruce |
| 6 | 1 | RP | Red Pine |
| 7 | 1 | SH | Unspecified Softwood |
| 8 | 1 | LT | Larch |
| 9 | 1 | WS | White Spruce |
| 11 | 2 | GOOD | Good |
| 12 | 2 | MEDIUM | Medium |
| 13 | 2 | POOR | Poor |
| 15 | 3 | D0 | No Crown Closure |
| 16 | 3 | D1 | 75% - > Crown Closure |
| 17 | 3 | D2 | 51% - 74% Crown Closure |
| 18 | 3 | D3 | 26% - 50% Crown Closure |
| 19 | 3 | NN | Null Value For The Non-forested Polyg |
| 21 | 4 | W | Working forest |
| 22 | 4 | R | Reserve Forest |
[21]:
sit.sit_data.disturbance_types
[21]:
| sit_disturbance_type_id | id | name | |
|---|---|---|---|
| 0 | 1 | DISTID1 | Natural forest fire |
| 1 | 2 | DISTID2 | Senescence |
| 2 | 3 | DISTID3 | Mountain pine beetle infestation |
| 3 | 4 | DISTID4 | ClearCut Harvesting |
[22]:
sit.sit_data.disturbance_events
[22]:
| Working_Species_Or_Leading_Species | Site_Quality | Density_Class | Working_Status | min_age | max_age | MinYearsSinceDist | MaxYearsSinceDist | LastDistTypeID | MinTotBiomassC | ... | MinSWMerchStemSnagC | MaxSWMerchStemSnagC | MinHWMerchStemSnagC | MaxHWMerchStemSnagC | efficiency | sort_type | target_type | target | disturbance_type | time_step | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | BF | GOOD | D1 | W | 80 | 200 | -1.0 | -1.0 | -1 | -1.0 | ... | -1.0 | -1.0 | -1.0 | -1.0 | 1.0 | SORT_BY_SW_AGE | Area | 200.0 | DISTID4 | 1 |
| 1 | BF | GOOD | D1 | W | 80 | 200 | -1.0 | -1.0 | -1 | -1.0 | ... | -1.0 | -1.0 | -1.0 | -1.0 | 1.0 | SORT_BY_SW_AGE | Area | 200.0 | DISTID4 | 2 |
| 2 | BF | GOOD | D1 | W | 80 | 200 | -1.0 | -1.0 | -1 | -1.0 | ... | -1.0 | -1.0 | -1.0 | -1.0 | 1.0 | SORT_BY_SW_AGE | Area | 200.0 | DISTID4 | 3 |
| 3 | BF | GOOD | D1 | W | 80 | 200 | -1.0 | -1.0 | -1 | -1.0 | ... | -1.0 | -1.0 | -1.0 | -1.0 | 1.0 | SORT_BY_SW_AGE | Area | 200.0 | DISTID4 | 4 |
| 4 | BF | GOOD | D1 | W | 80 | 200 | -1.0 | -1.0 | -1 | -1.0 | ... | -1.0 | -1.0 | -1.0 | -1.0 | 1.0 | SORT_BY_SW_AGE | Area | 200.0 | DISTID4 | 5 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 155 | BF | GOOD | D1 | W | 80 | 200 | -1.0 | -1.0 | -1 | -1.0 | ... | -1.0 | -1.0 | -1.0 | -1.0 | 1.0 | SORT_BY_SW_AGE | Area | 200.0 | DISTID4 | 156 |
| 156 | BF | GOOD | D1 | W | 80 | 200 | -1.0 | -1.0 | -1 | -1.0 | ... | -1.0 | -1.0 | -1.0 | -1.0 | 1.0 | SORT_BY_SW_AGE | Area | 200.0 | DISTID4 | 157 |
| 157 | BF | GOOD | D1 | W | 80 | 200 | -1.0 | -1.0 | -1 | -1.0 | ... | -1.0 | -1.0 | -1.0 | -1.0 | 1.0 | SORT_BY_SW_AGE | Area | 200.0 | DISTID4 | 158 |
| 158 | BF | GOOD | D1 | W | 80 | 200 | -1.0 | -1.0 | -1 | -1.0 | ... | -1.0 | -1.0 | -1.0 | -1.0 | 1.0 | SORT_BY_SW_AGE | Area | 200.0 | DISTID4 | 159 |
| 159 | BF | GOOD | D1 | W | 80 | 200 | -1.0 | -1.0 | -1 | -1.0 | ... | -1.0 | -1.0 | -1.0 | -1.0 | 1.0 | SORT_BY_SW_AGE | Area | 200.0 | DISTID4 | 160 |
160 rows × 33 columns
[23]:
sit.sit_data.yield_table
[23]:
| Working_Species_Or_Leading_Species | Site_Quality | Density_Class | Working_Status | leading_species | v0 | v1 | v2 | v3 | v4 | ... | v11 | v12 | v13 | v14 | v15 | v16 | v17 | v18 | v19 | v20 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | BF | GOOD | D1 | W | BF | 0.0 | 0.0 | 0.0 | 8.0 | 20.0 | ... | 112.0 | 113.0 | 113.0 | 113.0 | 113.0 | 113.0 | 113.0 | 113.0 | 113.0 | 113.0 |
1 rows × 26 columns
[24]:
print(json.dumps(sit.config, indent=4, sort_keys=True))
{
"import_config": {
"age_classes": {
"params": {
"path": "age_classes.csv"
},
"type": "csv"
},
"classifiers": {
"params": {
"path": "classifiers.csv"
},
"type": "csv"
},
"disturbance_types": {
"params": {
"path": "disturbance_types.csv"
},
"type": "csv"
},
"events": {
"params": {
"path": "disturbance_events.csv"
},
"type": "csv"
},
"inventory": {
"params": {
"path": "inventory.csv"
},
"type": "csv"
},
"transitions": {
"params": {
"path": "transition_rules.csv"
},
"type": "csv"
},
"yield": {
"params": {
"path": "growth_and_yield.csv"
},
"type": "csv"
}
},
"mapping_config": {
"disturbance_types": [
{
"default_dist_type": "Clearcut harvesting without salvage",
"user_dist_type": "ClearCut Harvesting"
},
{
"default_dist_type": "Wildfire",
"user_dist_type": "Natural forest fire"
},
{
"default_dist_type": "Stand\u2013replacing natural succession",
"user_dist_type": "Senescence"
},
{
"default_dist_type": "Insect disturbance",
"user_dist_type": "Mountain pine beetle infestation"
}
],
"nonforest": null,
"spatial_units": {
"admin_boundary": "Ontario",
"eco_boundary": "Boreal Shield East",
"mapping_mode": "SingleDefaultSpatialUnit"
},
"species": {
"species_classifier": "Working Species Or Leading Species",
"species_mapping": [
{
"default_species": "Balsam fir",
"user_species": "Balsam Fir"
},
{
"default_species": "Black spruce",
"user_species": "Black Spruce"
},
{
"default_species": "Western larch",
"user_species": "Western Larch"
},
{
"default_species": "Jack pine",
"user_species": "Jack Pine"
},
{
"default_species": "Other spruce",
"user_species": "Other Spruce"
},
{
"default_species": "Red pine",
"user_species": "Red Pine"
},
{
"default_species": "Unspecified softwood species",
"user_species": "Unspecified Softwood"
},
{
"default_species": "Tamarack/larch",
"user_species": "Larch"
},
{
"default_species": "White spruce",
"user_species": "White Spruce"
}
]
}
}
}
[ ]:
[ ]:
[ ]: